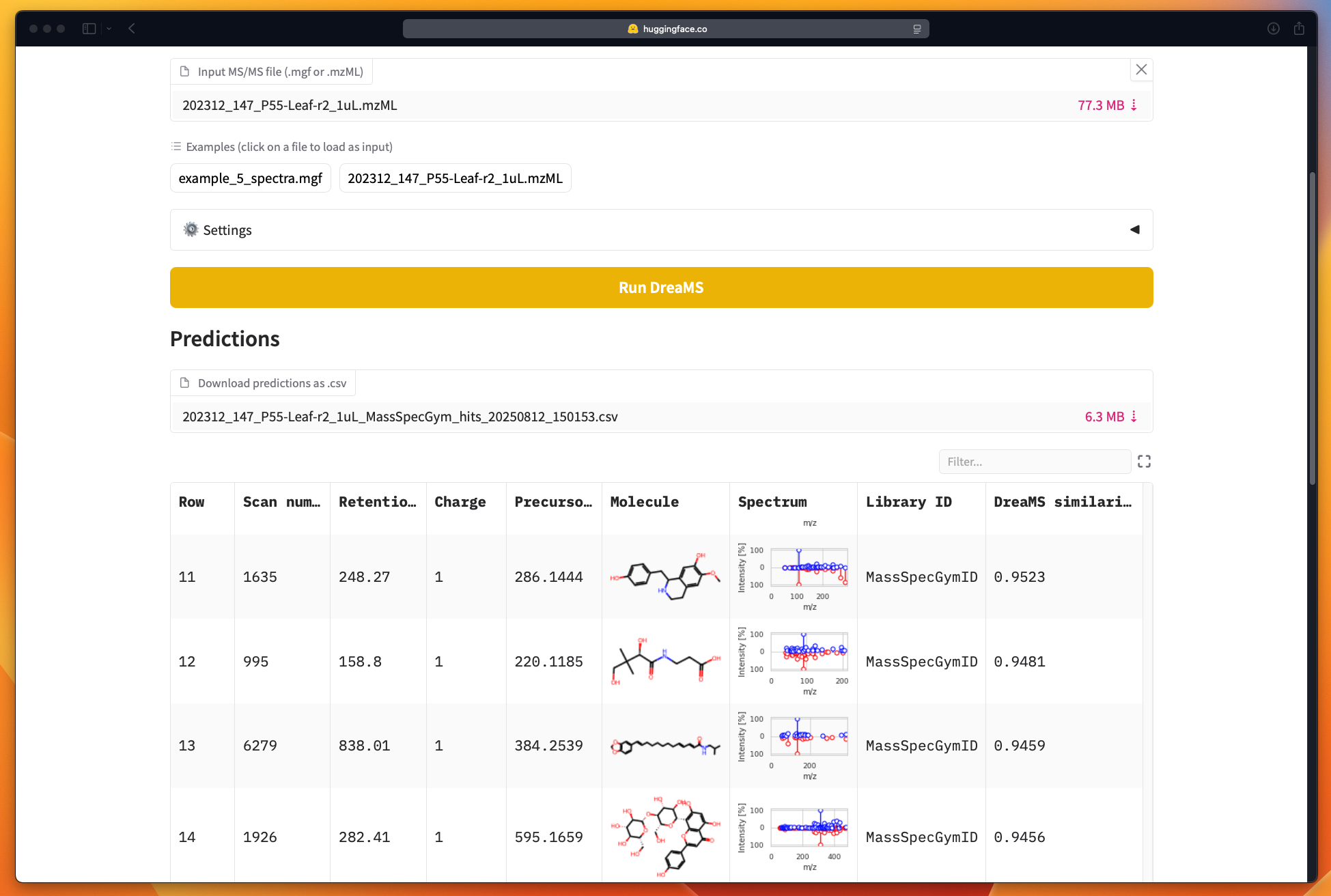
DreaMS is an innovative deep learning framework specifically engineered for the self-supervised learning of molecular representations from vast datasets of tandem mass spectra. Building upon millions of spectral observations, this tool extracts latent features that are highly informative about molecular structure and identity, thereby enabling advanced chemical analysis and characterization.
This powerful framework finds its application across a multitude of scientific domains, particularly within chemistry, computational chemistry, and various areas of life sciences requiring precise molecular identification and quantification. DreaMS is instrumental in advancing spectral analysis and characterization ecosystems, offering a robust solution for interpreting complex mass spectrometry data.
Practical applications and use cases for DreaMS are diverse and impactful. In metabolomics, it can be applied to problems such as the detection of specific beta-oxidation defects by analyzing acylcarnitine profiles, improving metabolite identification, and enhancing the confidence of metabolite annotation. For proteomics, DreaMS facilitates the identification of specific prey proteins in affinity purification-mass spectrometry (AP-MS) experiments and enhances the accuracy of protein identification in shotgun proteomics by generating more robust spectral representations. Furthermore, it addresses significant computational challenges in proteogenomics and peptide sequencing by providing an efficient means to process and interpret large-scale MS/MS data, allowing researchers to explore protein variants and post-translational modifications more effectively. By transforming raw mass spectral data into meaningful molecular representations, DreaMS accelerates discovery in areas ranging from biomarker identification in disease research to environmental monitoring and pharmaceutical development.
工具构建参数
| 主要语言 | Jupyter Notebook (98.28%) |
| 许可证 | MIT |